Protein conformational dynamics
and function
We are interested in understanding the dynamic-function paradigm in proteins.
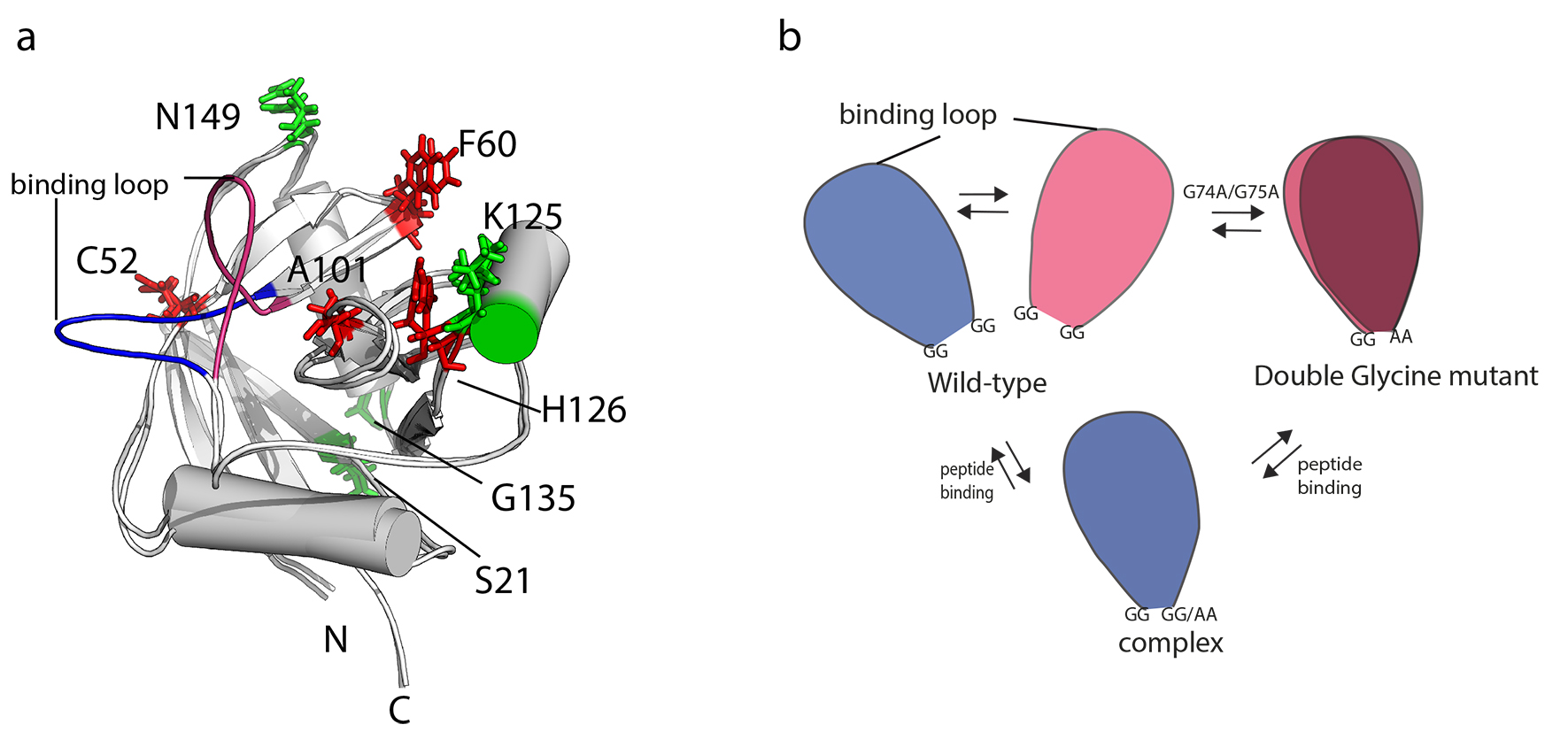
It is now generally accepted that the conformational dynamic landscape of proteins, which ranges from local perturbation to large structural rearrangements are relevant to protein function. The magnitude of this structural perturbation may cover a large range from 0.1-100 Å with a time frame from 10-12 seconds to > 1s. However, it is difficult to represent the spatial sampling of these motions. Several approaches are being developed to give a combined description of motional and functional propeties of proteins. One approach which is currently being developed in the our laboratory involves the determination of structural ensembles by the use of distances derrived from exact cross correlated relaxation (eNOE) and residual dipolar couplings. Application of this technique to the enzyme cyclophilin revelas that it exist in an open and closed state. (Link to movie)
A problem of solution NMR has been its limitation to small and medium size and isolated protein domains because sensitivity and resolution deteriorates with increasing size of marcromolecules. To solve this size issue and make solution NMR applicable to large marcromolecule, we are currently developing a tool for structral refinement for large protein and protein complexes by the use of eNOE distances derived from methyl-methyl cross relaxation. Beacuse of the experimental accuracy of the distances derived from eNOE it would be possible to used a subset of these distances as a ruler for calibrating and refining X-ray stuctures of large proteins and protein complexes. The eNOE distances could as well be use as benchmark for molecular dynamics simulations. We are currently looking at proteasome, membrane proteins and other larger complexes in this respect.